Student Researchers Investigate Cellular Stress Responses Using Novel Cell Imaging Techniques
Until recently, cells have been viewed in a static manner, which leaves out potentially crucial data and destroys the cells in the process. However, researchers in the Department of Chemical Engineering at the University of California, Davis, are working to combine live-cell imaging and image-based profiling can be used to study cellular stress responses in a more dynamic way and without killing the cells.

The novel approach of combining live-cell imaging — a technique that allows the researcher to image the cells and observe their response to external perturbations like viruses and drugs over a period of time — with image-based profiling to investigate autophagy was pioneered in UC Davis' Shah Lab by recent Ph.D. graduate Nitin Beesabathuni. Autophagy is a cell recycling process in which a cell breaks down unwanted or dysfunctional parts, keeping what it needs and discarding the rest; it's one of the body's ways of fighting off infection.
Beesabathuni found that he could use machine learning to look at thousands of features of the cell at once and identify patterns to determine which cells require further investigation, and hopes that these methods can also be applied to other cellular research, including cancer and viruses, leading to new treatments.
"As humans, we can look at a couple or a handful of features, but we can't look at thousands of features at once," he said. "But using live-cell imaging, machine learning and image-based profiling, you can look at how the cell changes every 30 minutes after any perturbation like viruses or drug treatment."
A New Way of Looking at Things
Beesabathuni, who finished his Ph.D. in chemical engineering at UC Davis in March 2023, was among the first graduate students to work in the Shah Lab, which focuses on investigating viruses and how they convert host cells into microscopic virus factories and is led by Assistant Professor Priya Shah, who holds appointments in chemical engineering and microbiology and molecular genetics. When Beesabathuni was assigned to research autophagy and viruses, he realized that people looking at autophagy were largely using static measurements.
"I came up with this idea of using the basic principles of chemical engineering to measure the kinetics of these complicated cellular processes. It's a new way of looking at things and it's more comprehensive and systematic," said Beesabathuni.
The lab set up a live-cell imaging workflow, a particular boon to their research on autophagy because being able to observe the initial phase of the cells' immediate response to a drug can be critical for determining new treatments.
Image of Success
Beesabathuni also had the issue of sampling the cells in an efficient way: He was using the destructive sampling approach, a time-consuming and tedious method in which a cell can only be sampled once before it is destroyed.
When considering testing drugs on an industrial scale, this was a non-starter for Beesabathuni. But a lightbulb moment came during a lighthearted game with a fellow Ph.D. student.
"I would show him images and ask him to guess which treatment it was, like was it an inducer or an inhibitor, and he used to guess quite accurately," he said. "It got me thinking that instead of doing this very complicated process of measuring kinetics of autophagy by destructive sampling, can we use machine learning where we can look at the images of the cells and comprehend what is an inducer and inhibitor without destructive sampling?"

At that point, Eshan Thilakaratne joined the Shah Lab as an undergraduate student researcher in chemical engineering. Thilakaratne's task was to write the foundational codes for a computational pipeline in which they could profile the cells using imaging (hence, image-based profiling). He investigated different features to see how he could implement them in the coding language Python and whether the features could be used to extract useful information from cells.
Beesabathuni compares image-based profiling to a social media algorithm, but for screening drugs on an industrial scale instead of finding reels on Instagram.
"Let's say, for example, I'm on social media and I love soccer, and I like soccer videos and I look at those videos longer. The algorithms measure all these things, and compares it with the reference data and says, 'Oh, they must like soccer, so let's give them more soccer,'" he said. "We are doing exactly the same thing, except with drugs. Let's say there's a drug that can treat the disease and we know that, and we find more drugs like that. We're looking at whether the cells look more similar to the drug that we know treats disease."
So, Beesabathuni and Thilakaratne built the computational pipeline for applying image-based profiling to autophagy from scratch, which has been presented with Shah in a pre-print, or a non-peer reviewed work in progress. While the Shah Lab is using the pipeline to investigate autophagy responses to chemical treatment, the group is optimistic about its other potential applications, like researching chemical treatments for cancer cells or viral replication in virus-infected cells.
A Foundation to be Built Upon
Beesabathuni and Thilakaratne have since moved on from UC Davis — Beesabathuni is using image-based profiling in his drug research at Cambridge-based pharmaceutical company Merck and Thilakaratne is now doing genomic surveillance on COVID strains for the California Department of Public Health. However, researchers in Shah Lab like Neil Adia, a second year Ph.D. student in chemical engineering, are using these lab-developed tools as a foundation for their own research.
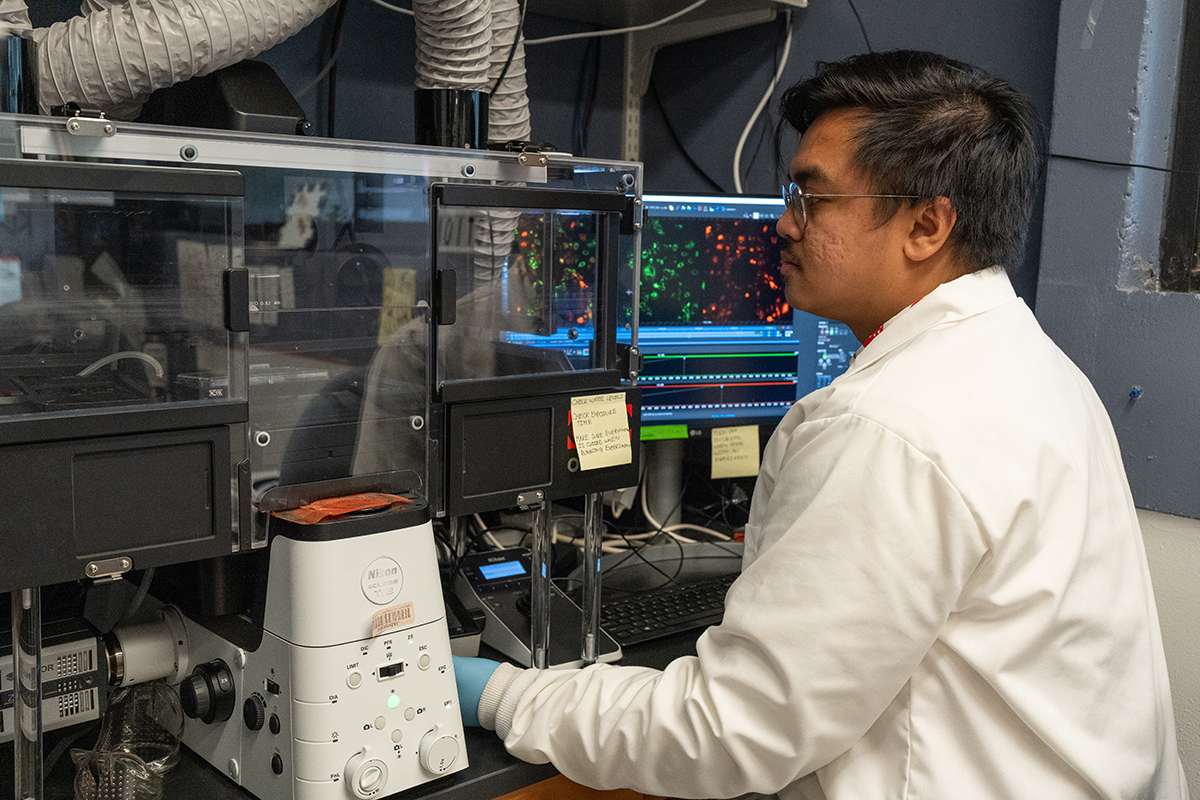

Adia is investigating the interplay between Zika virus and autophagy. When he began using live-cell imaging and image-based profiling, he didn't realize the full potential of this set of tools at his disposal for extracting information from cell images.
"When I first joined the Shah Lab, I was under the assumption, 'Ok, we're going to infect cells, and we're going to see the different fluorescent proteins that are there.' And that's it," he said. "It took me a while to realize that there's a lot more information within the images in terms of how things look and in all these features that we extract."
Adia states that live-cell imaging and image-based profiling has a profound impact on his work. Not only do these techniques allow for less tedium in the actual experimental process than the more traditional destructive sampling method, but they create a clearer, more holistic view of how infections cause cells' morphology to change over time.
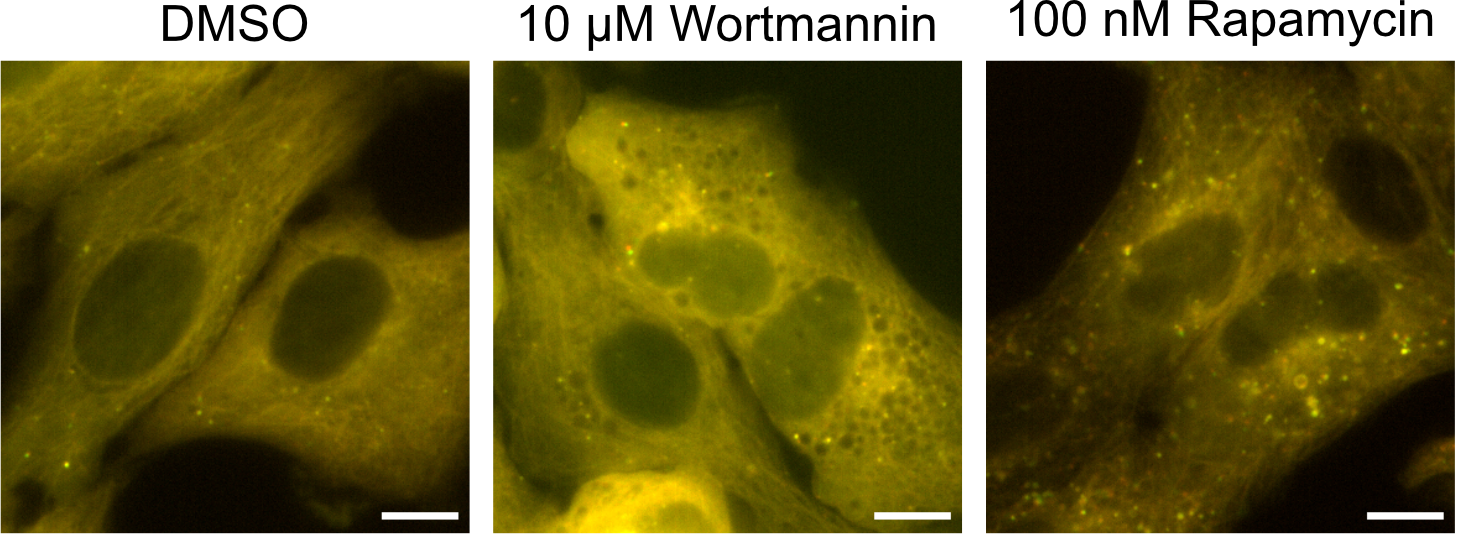
Adia plans to use the base of what Beesabathuni and Thilakaratne built and add features to personalize the process to his needs in terms of Zika virus and autophagy and maximize his workflow. The two alumni agree that the fact that these tools are foundational and can be built upon to be used in many different research areas – drugs, cancer, neurodegeneration, etc. — is one of the most exciting things about their potential.
"It's really cool that the research and this project that Nitin started really planted the seed for the lab, in terms of where the research is going to go," said Thilakaratne. "I think there's going to be a lot of branches to it."
